what must be true about the inner mitochondrial membrane for the gradient to work
- Review
- Open Access
- Published:
Structure and function of mitochondrial membrane protein complexes
BMC Biology volume thirteen, Commodity number:89 (2015) Cite this article
Abstruse
Biological energy conversion in mitochondria is carried out by the membrane protein complexes of the respiratory chain and the mitochondrial ATP synthase in the inner membrane cristae. Recent advances in electron cryomicroscopy take made possible new insights into the structural and functional system of these complexes in the membrane, and how they modify with age. This review places these advances in the context of what is already known, and discusses the fundamental questions that remain open up but can at present exist approached.
Mitochondria are the powerhouses of the prison cell. In all eukaryotes that practice not depend on photosynthesis, the mitochondria are the main source of adenosine triphosphate (ATP), the energy-rich compound that drives key jail cell functions. These functions include force generation (for example, in muscle contraction and cell partition), the biosynthesis, folding and deposition of proteins, and the generation and maintenance of membrane potentials. ATP is produced on a massive scale in the man body, amounting to fifty kg per day in a healthy adult, but considerably more in a long-distance runner. ATP is generated by the mitochondrial ATP synthase from ADP and phosphate ions. These are the products of ATP hydrolysis at the sites where energy is needed in the cell. Autonomously from cellular respiration and ATP synthesis, mitochondria have numerous other essential functions, including the production of NADH and GTP in the citric acid cycle, the biosynthesis of amino acids, heme groups and atomic number 26-sulfur clusters or the synthesis of phospholipids for membrane biogenesis. They also act in calcium signaling [1], stress responses [2] and generally as cellular signaling hubs [3]. Not surprisingly, mitochondria play a key role in human health. Mitochondrial dysfunction is the cause of astringent, often maternally inherited diseases. Moreover, mitochondria are deeply implicated in apoptosis and ageing [4].
In many respects, mitochondria resemble α-proteobacteria, from which they are thought to take originated by endocytosis some 1.half dozen billion years ago. The most striking evidence of this evolutionary human relationship is the shut homology of bacterial and mitochondrial respiratory chain complexes. Mitochondria have their ain genetic system, which uses a distinct DNA lawmaking that differs both from that of their bacterial ancestors and their eukaryotic hosts [5]. They accept their own protein translation machinery, complete with ribosomes, tRNAs and associated protein factors that more or less resemble those of their bacterial ancestors. Very recently, the start high-resolution structure of a mitochondrial ribosome, determined by single-particle electron cryomicroscopy (cryo-EM), has revealed a fascinating patchwork of similarities to and differences from bacterial ribosomes [vi]. Nevertheless, mitochondria make surprisingly little apply of their specialized protein production machinery. In the class of evolution they have transferred up to 99 % of their genes to the nucleus. Today, the vast majority of mitochondrial proteins are produced in the cytoplasm and imported into the organelle by an elaborate set of protein translocases [7]. In humans, but 13 mitochondrial proteins are organelle-encoded, all of them cardinal, hydrophobic subunits of respiratory chain complexes or of the ATP synthase.
Mitochondria are highly dynamic [eight]. In the cell, they grade a tubular network that constantly changes by partition and fusion (Boosted file i). Both processes are accomplished by multi-component molecular machineries that include a number of dynamin-related GTPases [nine, 10]. When mitochondria are isolated from cells, the network breaks up into fragments that spontaneously reseal. Isolated mitochondria are fully competent for respiration and ATP synthesis [xi]. They maintain their membrane limerick, organization and membrane potential, as well every bit the power to fuse [12] and to import proteins [7]. We owe much of what we know most mitochondria and how they work at the molecular level to in vitro studies with isolated mitochondria, or fifty-fifty mitochondrial membrane fractions, which still comport out oxidative phosphorylation and ATP synthesis [13].
Mitochondria can be seen in the light microscope, but their detailed internal construction is simply revealed past electron microscopy. In the 1990s, the structure of mitochondria was investigated past electron tomography of thin plastic sections [14]. While this yielded striking three-dimensional (3D) images of their internal membrane system, molecular detail was lost due to chemic fixation, dehydration and heavy-metal staining. Cryo-EM of unfixed, unstained organelles is now revealing the architecture of mitochondrial membranes and their macromolecular components at increasing levels of detail. Single-particle cryo-EM of isolated, detergent-solubilized membrane poly peptide complexes reaches near-diminutive resolution [15, 16]. Electron cryo-tomography (cryo-ET) of intact isolated mitochondria or mitochondrial membranes is resolving their macromolecular components in situ [17], and averaging of tomographic volumes can attain sub-nanometer resolution [18].
Mitochondrial membranes and membrane compartments
As ubiquitous, semi-autonomous cellular organelles, mitochondria are separated from the cytoplasm by the outer and inner mitochondrial membrane (Fig. ane). The outer membrane is porous and freely traversed by ions and small, uncharged molecules through pore-forming membrane proteins (porins), such as the voltage-dependent anion channel VDAC [nineteen]. Whatever larger molecules, especially proteins, have to exist imported by special translocases. Because of its porosity, there is no membrane potential across the outer membrane. By contrast, the inner membrane is a tight diffusion barrier to all ions and molecules. These tin simply get across with the aid of specific membrane transport proteins, each of which is selective for a particular ion or molecule. As a issue of its ion selectivity, an electrochemical membrane potential of almost 180 mV builds up across the inner mitochondrial membrane. The inner membrane is where oxidative phosphorylation takes place in a suite of membrane protein complexes that create the electrochemical gradient across the inner membrane, or apply it for ATP synthesis.
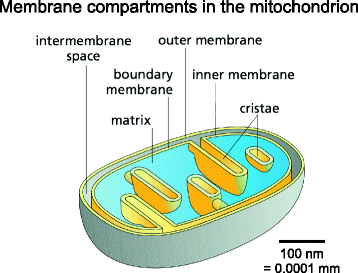
Membrane compartments in the mitochondrion. The outer membrane separates mitochondria from the cytoplasm. Information technology surrounds the inner membrane, which separates the inter-membrane space from the poly peptide-dense central matrix. The inner membrane is differentiated into the inner boundary membrane and the cristae. The 2 regions are continuous at the crista junctions. The cristae extend more or less deeply into the matrix and are the master sites of mitochondrial energy conversion. The shallow proton gradient between the inter-membrane space (pH seven.ii–7.4) and the matrix (pH 7.nine–viii) drives ATP production by the ATP synthase in the membranes of the cristae. (Adapted from Effigy 14–8 C in Alberts B. et al. Molecular Biology of the Cell. half-dozenth ed. New York: Garland Science; 2014, with permission of the publisher [© 2015 from Molecular Biology of the Jail cell, 6th Edition past Alberts et al. Reproduced by permission of Garland Science/Taylor & Francis Group LLC])
The inner and outer membranes of mitochondria ascertain three compartments within the organelle, each with its distinct office and corresponding protein components. The innermost compartment, surrounded by the inner membrane, is the mitochondrial matrix. It is the equivalent of the bacterial cytoplasm, from which it is distinguished by a pH of 7.9 to viii [20], similar to that in the chloroplast stroma. The high pH of the mitochondrial matrix creates the trans-membrane electrochemical gradient that drives ATP synthesis (come across below). The mitochondrial matrix is the site of organellar Deoxyribonucleic acid replication, transcription, poly peptide biosynthesis and numerous enzymatic reactions. Mitochondrial Dna is compacted by the mitochondrial transcription factor TFAM into supramolecular assemblies chosen nucleoids, of which there are about 1000 per cell [21]. Nucleoids are roughly spherical, with a diameter of ~100 nm, each containing 1 re-create of mitochondrial DNA [22]. Mitochondrial ribosomes are membrane-fastened, as their simply products (in man cells) are hydrophobic membrane poly peptide subunits, which integrate directly into the inner membrane upon translation. A ~25 Å resolution structure of the membrane-bound mitochondrial ribosome has recently been obtained past cryo-ET and sub-tomogram averaging [23].
The biosynthetic reactions that happen in the matrix include those of the citric acid cycle. Equally each reaction is catalyzed past its specific enzyme, the mitochondrial matrix has a high protein density of up to 500 mg/ml, shut to that in a protein crystal. For cryo-ET of intact organelles, the high matrix density has the disadvantage of obscuring internal detail.
The equivalent of the periplasm in the bacterial ancestors of mitochondria is the intermembrane space. This is the ~twenty nm gap between the outer membrane and the function of the inner membrane that is known every bit the inner boundary membrane. All matrix proteins imported into the mitochondrion from the cytoplasm must pass through the outer and inner membrane and therefore also through the intermembrane infinite. Conventional EM of thin plastic sections suggested sites of direct contact between the lipid bilayers of the inner and outer membrane [24, 25], merely these seem to exist artifacts of fixation and dehydration. Protein translocases of the outer (TOM) and inner (TIM) membrane grade a supercomplex that has been visualized by cryo-ET [26]. The TOM/TIM supercomplex spans the intermembrane space and appears to be held together by the polypeptide in transit. The inner purlieus membrane must contain large numbers of the carrier proteins that shuttle ions, ATP, ADP and pocket-size metabolites betwixt the cytoplasm and the matrix. These small membrane proteins include most notably the 33 kDa ATP/ADP carrier [27], likewise as numerous other related and unrelated membrane transporters.
The inner membrane forms invaginations, called cristae, that extend deeply into the matrix. The cristae ascertain the third mitochondrial compartment, the crista lumen. The crista membranes contain most, if not all, of the fully assembled complexes of the electron transport chain and the ATP synthase (Fig. 2). The crista lumen contains large amounts of the pocket-sized soluble electron carrier protein cytochrome c. The mitochondrial cristae are thus the main site of biological energy conversion in all non-photosynthetic eukaryotes.
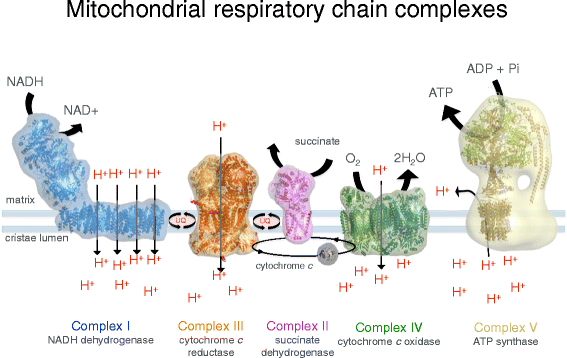
Membrane protein complexes of the respiratory chain. Electron transport complexes I (NADH/ubiquinone oxidoreductase, blue), II (succinate dehydrogenase, pinkish), III (cytochrome c reductase, orange), IV (cytochrome c oxidase, greenish) and the mitochondrial ATP synthase (likewise known as complex V, tan) work together in oxidative phosphorylation to harness free energy for the cell. Complexes I, III and Four pump protons across the cristae membrane, creating the proton slope that drives ATP synthesis. UQ ubiquinol. (Adapted from Davies KA, Daum B. Biochem. Soc. Trans. 2013;41:1227–34.)
The inner membrane cristae
Cristae were start discovered by electron microscopy in thin sections of plastic-embedded cells and tissues [28, 29]. They are deejay-like lamellar, tubular or pocketbook-similar extensions of the inner purlieus membrane, and are continuous with information technology at the crista junctions. Crista junctions in mitochondria of post-mitotic heart or liver cells are modest round apertures of ~25 nm diameter [14, 30]. In cells that divide frequently, such as yeasts, the crista junctions tend to form 25 nm slits in the purlieus membrane that are up to a few 100 nm long [26, thirty]. In mitochondria of all organisms, the mitochondrial contact site and cristae organizing (MICOS) system [31], an assembly of ane soluble and five membrane proteins, anchors the cristae to the outer membrane. Cristae of yeast strains where particular MICOS components take been knocked out await like concentric onion rings and take few if any junctions [32].
Information technology is thought that the MICOS complex forms a diffusion barrier within the inner membrane at the crista junctions to account for the apparent lateral segregation of proteins between the cristae and boundary membranes. Testify of such differences comes primarily from electron microscopy, because it has not been possible to dissever cristae and boundary membranes biochemically. Immuno-staining of thin plastic sections has shown that respiratory chain complexes reside in the cristae rather than in the boundary membrane, whereas components of the protein translocases are found in the purlieus and outer membranes [33]. Cryo-ET of intact mitochondria, which resolves large membrane protein complexes in situ, did not reveal any such assemblies (for example, the ATP synthase or complex I) in the boundary membrane [17], suggesting that the protein complexes of the inner membrane are indeed laterally segregated.
In tissues with a high free energy demand, such as skeletal or heart muscle, the cristae are closely stacked disk-like lamellae that have up most of the mitochondrial book (Fig. 3). In animal tissues with lower energy demand, such as liver or kidney, the cristae are less closely stacked, leaving more room for the matrix with its biosynthetic enzymes. In all mitochondria, the cristae business relationship for most of the inner membrane surface, highlighting their importance for cellular physiology.

Tomographic volume of mouse center mitochondrion. a Three-dimensional volume of a mouse heart mitochondrion adamant past cryo-ET. The outer membrane (grey) envelops the inner membrane (light blue). The inner membrane is highly folded into lamellar cristae, which criss-cross the matrix. b Tomographic slice through the map volume. The dumbo matrix, which contains most of the mitochondrial poly peptide, appears dark in the electron microscope, whereas the intermembrane infinite and crista lumen appear lite because of their lower poly peptide content. The inner boundary membrane follows the outer membrane closely at a distance of ~twenty nm. The inner membrane turns sharply at the crista junctions, where the cristae join the inner boundary membrane. (Courtesy of Tobias Brandt)
ATP synthase forms rows of dimers in crista membranes
The mitochondrial Fi-Fo ATP synthase is the most conspicuous protein circuitous in the cristae. The ATP synthase is an ancient nanomachine that uses the electrochemical proton slope across the inner mitochondrial membrane to produce ATP by rotatory catalysis [34]. Protons moving through the Fo circuitous in the membrane drive a rotor ring composed of 8 (in mammals [35]) or ten (in yeast [36]) c-subunits. The key stalk propagates the c-ring torque to the catalytic F1 head, where ATP is generated from ADP and phosphate through a sequence of conformational changes. The peripheral stalk prevents unproductive rotation of the Fi head against the Fo circuitous.
For many years it was causeless that the ATP synthase and other energy-converting complexes are randomly distributed over the inner membrane. The first hint that this is not the case came from deep-etch freeze-fracture electron microscopy, which revealed double rows of macromolecular complexes in the tubular cristae of the single-cell ciliate Paramecium [37]. The double rows were thought to be linear arrays of mitochondrial ATP synthase. This is indeed what they are, but information technology could but be shown unambiguously more than than twenty years subsequently by cryo-ET [30, 38], which revealed rows of ATP synthase dimers in mitochondria of all species investigated [30] (Fig. four). Until so, the rows were thought to be peculiar to Paramecium.
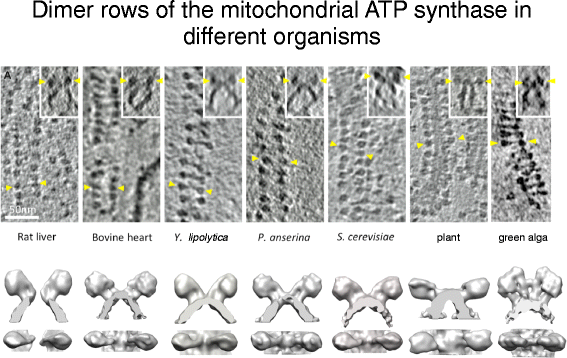
Double rows of ATP synthase in seven different species. Peak row: slices of 3D volumes obtained past cryo-ET with rows of ATP synthase dimers. Insets prove side views of the dimers in the membrane. Yellowish arrowheads indicate F1 heads of one dimer. Calibration bar = 50 nm. Bottom row: Surface representations of subtomogram averages. (Adapted from [17])
The linear arrays of ATP synthase dimers are a ubiquitous and fundamental aspect of all mitochondria. They are e'er found along the most tightly curved regions along the crista ridges (Additional file 2), or effectually narrow tubular cristae. Subtomogram averages indicate that dimers from fungi and mammals are indistinguishable at low resolution, whereas those from plants, algae and protists differ in dimer angle or position of the peripheral stalk relative to the catalytic Fone head (Fig. four). However, the basic assembly of ATP synthase complexes into dimers and their association into long rows forth the crista ridges are conserved. Coarse-grained molecular dynamics simulations indicate that the dimers bend the lipid bilayer and, as a issue, cocky-acquaintance into rows [thirty]. Accordingly, row formation does not crave specific protein interactions, but is driven by the energy of elastic membrane deformation. Cryo-ET of isolated, detergent-solubilized ATP synthase dimers reconstituted into proteoliposomes has provided experimental evidence that the dimers do in fact self-associate into rows and that they induce a local curvature in the lipid bilayer (Blum, Davies, Kühlbrandt, unpublished results). Therefore, it is in fact the dimers that curve the membrane, not the other way round.
The structure of a mitochondrial ATP synthase dimer has now been determined past single-particle cryo-EM at six–7 Å resolution [39]. Surprisingly, the structure revealed a hairpin of long, membrane-intrinsic α-helices in the a-subunit next to the Fo rotor ring, also equally aqueous half-channels on either side of the membrane for proton translocation. The long helices appear to play a central role in this process, as they are preserved in all rotary ATPases [40]. Together with the fitted high-resolution X-ray structures of the catalytic Fane head and the rotor band in the membrane, the cryo-EM map provides the first complete moving picture of this pivotal mitochondrial membrane complex (Fig. 5) and goes a long way towards explaining its mechanism [40].
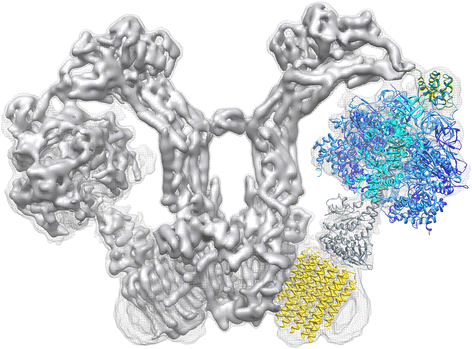
Structure of the mitochondrial ATP synthase dimer from Polytomella sp. Side view of the two mitochondrial ATP synthase in the Five-shaped dimer. Ane protomer is fitted with atomic models PDB 2WSS [63] (α-subunits, cyan; β-subunits, blueish; γδε subunits, grayness; OSCP, dark-green) and PDB 3U2Y [64] (yellow, c 10-ring). Density threshold levels are 1σ (mesh) or 7σ (solid surface). (Adapted from [40])
The ubiquitous nature of the dimer rows raises the question as to the biological significance of this striking, conserved organisation. In yeast the two protomers are linked past the dimer-specific ATP synthase subunits e and g. If either subunit is knocked out, just monomeric ATP synthase is found in the inner membrane [30] and the usual lamellar or tubular cristae practise not grade [30, 41]. ATP synthase dimers and dimer rows are thus a prerequisite for proper cristae formation. Although the loss of the dimer-specific subunits is non lethal, it is a serious disadvantage. Yeast subunit e and yard knockout strains grow up to 60 % more slowly than wild type and their mitochondria but have about half the membrane potential [42]. This raises the farther question about the role of the cristae, and hence the dimer rows, in cellular physiology and fitness. Most likely the invaginations forbid protons that are pumped into the crista lumen by the respiratory chain from escaping speedily to the inter-membrane space and the cytoplasm, so that they tin can be harnessed more efficiently for ATP product. In this style, the cristae, and hence the dimer rows, would contribute to effective ATP synthesis.
Prokaryotic ATP synthases lack the dimer-specific subunits, and ATP synthase dimers or dimer rows take not been found in bacterial or archaeal inner membranes, which too do not have cristae. The cristae and dimer rows may thus exist an adaptation that enables mitochondria to satisfy the high energy need of eukaryotic cells with the bachelor, shallow proton slope of effectually 0.half-dozen–0.8 pH units. ATP synthase dimers have recently been implicated in the germination of the permeability transition pore [43] that triggers apoptosis. On the ground of the structure of the mitochondrial ATP synthase dimer [39] or the dimer rows [30], however, information technology is difficult to run into how they might form a membrane pore.
Respiratory chain complexes and supercomplexes
The proton gradient across the cristae membrane is generated by three large membrane poly peptide complexes of the respiratory chain in the cristae, known as complex I (NADH/ubiquinone oxidoreductase), III (cytochrome c reductase) and Iv (cytochrome c oxidase) (Fig. 2). Complex I feeds electrons from the soluble carrier molecule NADH into the respiratory chain and transfers them to a quinol in the membrane. The free energy released in the electron transfer reaction is utilized for pumping iv protons from the matrix into the crista lumen. Complex Iii takes the electrons from the reduced quinol and transfers them to the pocket-size, soluble electron carrier protein cytochrome c, pumping i proton in the procedure. Finally, complex Iv transfers the electrons from cytochrome c to molecular oxygen and contributes to the proton gradient by using up four protons per consumed oxygen molecule to make h2o. Complex II (succinate dehydrogenase) transfers electrons from succinate directly to quinol and does not contribute to the proton gradient.
The respiratory chain complexes accept been studied in bully item for decades. High-resolution X-ray structures are available for mitochondrial complex III [44] and IV [45]. At a molecular mass of ~1 megadalton (MDa), mitochondrial complex I is far larger and has more than subunits than complexes III and Four put together. As yet at that place is no Ten-ray construction of the mammalian complex, but very recently a ~iii.vi Å X-ray structure of complex I from the obligate aerobic yeast Yarrowia lipolytica has been obtained [46]. Comparison to the loftier-resolution X-ray construction of the ~550 kDa complex I from the thermophilic bacterium Thermus thermophilus [47] indicates that the 14 conserved core subunits have substantially the same structure in both, including iii proton antiporter modules in the membrane and eight iron-sulfur clusters in the matrix arm. The mitochondrial complex has about three times as many protein subunits as its bacterial ancestor. Nearly functions of the extra subunits are unknown, but many of them are likely to work in assembly or the regulation of complex I function. Features that are conserved from leaner to mitochondria include a long horizontal α-helix on the matrix side that may stabilize the membrane domain. The recent 5 Å single-particle cryo-EM structure of bovine heart complex I (Fig. 6) has resolved the proton-translocating modules, iron-sulfur clusters and long horizontal helix, and 14 of the 31 supernumerary mammalian complex I subunits have been identified [48]. Still, the way in which electron transfer from NADH to ubiquinone in complex I is coupled to proton translocation is still unknown, and much else remains to be discovered.
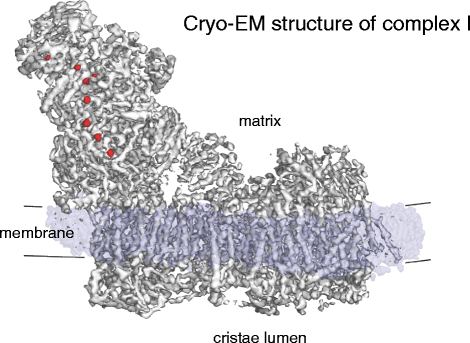
Cryo-EM structure of bovine eye complex I. Mitochondrial complex I (~1 MDa) has a matrix arm and a membrane arm. The matrix arm contains a row of eight iron-sulfur clusters (ruby-red) that conduct electrons from NADH to ubiquinol at the junction of the matrix and membrane arms (Fig. 7). The membrane arm consists of 78 trans-membrane helices, including iii proton-pumping modules. (Adapted from [51]; EMDB lawmaking 2676)
Not dissimilar the ATP synthase, which forms dimer rows in the cristae, the proton pumps of the electron transport chain assemble into supercomplexes or 'respirasomes'. Respiratory concatenation supercomplexes were first postulated on the footing of blue-native gels of yeast and bovine heart mitochondria solubilized in the balmy detergent digitonin [49]. Negative-stain electron microscopy [50] and single-particle cryo-EM [51] of the ane.7 MDa bovine eye supercomplex revealed that it consists of one re-create of complex I, one complex III dimer, and one complex IV monomer. 10-ray structures of the component complexes were fitted to the 3D map (Fig. seven) [51], indicating the path of the electron from NADH via the iron-sulfur clusters of complex I and ubiquinol to the prosthetic groups of complex Iii, and finally to molecular oxygen in complex 4. Genetic bear witness provides strong support for the being of respirasomes in vivo [52], simply they were long thought to be artifacts of detergent solubilization, notwithstanding their well divers structure. Recent cryo-ET piece of work has shown that they practise exist in cristae membranes of bovine heart mitochondria (Davies and Kühlbrandt, unpublished results). Saccharomyces cerevisiae, which lacks complex I, still has a respiratory concatenation supercomplex consisting of complex III and IV [53]. Far from beingness randomly distributed in the membrane, the ATP synthase and electron transport complexes of the respiratory chain thus grade supramolecular assemblies in the cristae, in a way that is essentially conserved from yeast to humans (Fig. 8). A clear functional office of mitochondrial supercomplexes has non yet been established. They may make electron transfer to and from ubiquinone in complexes I and III more efficient, as the relative positions and orientations of the two complexes are precisely aligned rather than random. However, there is no direct evidence that this makes a difference. The supercomplexes may simply help to avert random, unfavorable poly peptide–poly peptide interactions in the packed environment of the inner mitochondrial membrane [54]. Alternatively, they may control the ratio of respiratory chain complexes in the membrane, or aid their long-term stability.
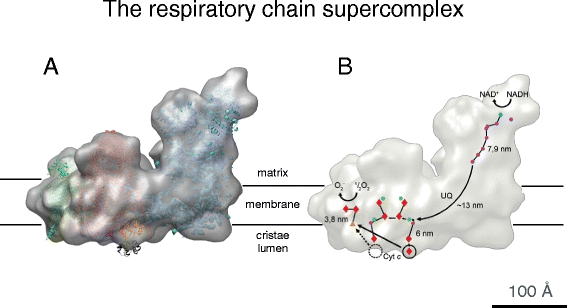
Cryo-EM structure of the i.seven MDa bovine heart respiratory concatenation supercomplex. a The supercomplex consists of ane copy of NADH dehydrogenase (circuitous I, blueish), a cytochrome b-c one dimer (complex 3, pinkish), and a single copy of cytochrome c oxidase (complex Iv, dark-green). b The ubiquinol (UQ) binding sites of complexes I and III and the curt distance betwixt the cytochrome c binding sites in complexes III and Four, which would favor efficient electron transfer. Cofactors active in electron transport are marked in yellow (FMN), orange (fe–sulfur clusters), dark blue (quinols), red (hemes), and green (copper atoms). Arrows indicate the electron path through the supercomplex. (Adapted from [51])
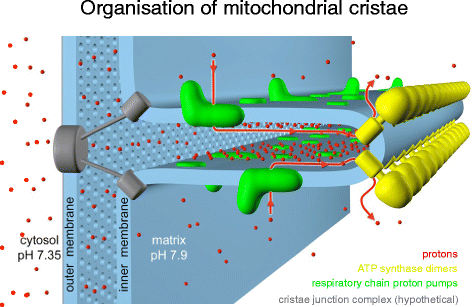
ATP synthase dimer rows shape the mitochondrial cristae. At the cristae ridges, the ATP synthases (yellow) form a sink for protons (scarlet), while the proton pumps of the electron send concatenation (green) are located in the membrane regions on either side of the dimer rows. Guiding the protons from their source to the proton sink at the ATP synthase, the cristae may piece of work as proton conduits that enable efficient ATP product with the shallow pH gradient between cytosol and matrix. Crimson arrows show the direction of the proton menstruum. (Adapted from [17])
A master protein component of the crista lumen is the small soluble electron carrier protein cytochrome c that shuttles electrons from complex III to complex Iv. If released into the cytoplasm, cytochrome c triggers apoptosis [55]. It is imperative, therefore, that cytochrome c does non leak from the cristae and that the outer membrane remains tightly sealed during mitochondrial fission and fusion.
Membrane rearrangement during cellular ageing
Ageing is a cardinal yet poorly understood biological process that affects all eukaryotic life. Deterioration in mitochondria is clearly seen in ageing, only details of the underlying molecular events are largely unknown. Cryo-ET of mitochondria from the short-lived (~18 days) model organism Podospora anserina revealed profound historic period-dependent changes in the inner membrane architecture [56]. In normal mitochondria of young cells, the cristae beetle deeply into the matrix. Formation of cristae depends both on the rows of ATP synthase dimers along the edges [30] and on the MICOS complex at the crista junctions [32]. With increasing age, the cristae recede into the inner boundary membrane and the inter-membrane space widens. The MICOS complex most likely has to come apart for this to happen. Eventually, the matrix breaks upward into spherical vesicles within the outer membrane. The ATP synthase dimer rows disperse and the dimers dissociate into monomers. As the inner membrane vesiculates, the sharp local curvature at the dimer rows inverts, then that the ATP synthase monomers are surrounded by a shallow concave membrane environment, rather than the sharply convex curvature at the crista ridges (Fig. 9). Finally, the outer membrane ruptures, releasing the inner membrane vesicles, along with apoptogenic cytochrome c, into the cytoplasm. Cytochrome c activates a cascade of proteolytic caspases, which degrade cellular proteins [55]. The prison cell enters into apoptosis and dies.
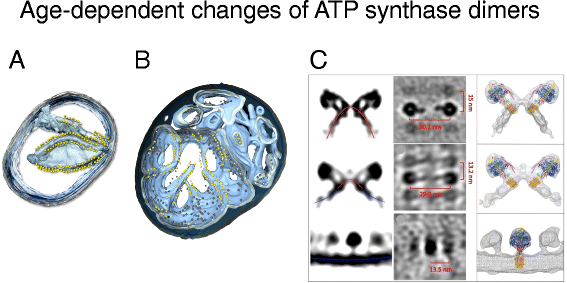
Changes of inner membrane morphology and ATP synthase dimers in ageing mitochondria. Tomographic volumes of mitochondria isolated from young (six-day-quondam) (a) and ageing (13-twenty-four hour period-former) (b) cultures of the model organism Podospora anserina. In young mitochondria, the ATP synthase dimers are bundled in rows along highly curved inner membrane ridges (Movie S2). In ageing mitochondria, the cristae recede into the boundary membrane, with ATP synthases dimer rows along the shallow inner membrane ridges. Outer membrane, transparent grey; inner membrane, calorie-free blue. ATP synthase F1 heads are shown as yellowish spheres. c Side and height views (left and centre) of subtomogram-averaged ATP synthase dimers from 6-twenty-four hours-old (acme row), 9-day-sometime (centre), and 15-day-old P. anserina mitochondria. Right: subtomogram averages with fitted Ten-ray models. Crimson lines, convex membrane curvature (as seen from the matrix); blueish lines, concave membrane curvature. (Adjusted from [56])
The observed morphological changes during ageing in P. anserina mitochondria may be a direct or indirect consequence of cumulative oxidative harm, which is known to promote cellular and organismal senescence [57]. The electron-transfer reactions in complexes I and Three generate reactive superoxide radicals equally side products [58], which crusade damage to mitochondrial proteins and DNA, as well as to other cellular components. Senescent mitochondria that lack cristae and ATP synthase dimers would non exist able to provide sufficient ATP to maintain essential cellular functions. Cells unremarkably bargain with oxidative damage by oxygen radical scavenging enzymes such as superoxide dismutase or catalase, likewise as past mitochondrial fission and fusion. Damaged or dysfunctional mitochondria are either complemented with an undamaged part of the mitochondrial network by fusion or sorted out for mitophagy [59]. During ageing, fission overpowers fusion and the mitochondrial network fragments [60]. This prevents the complementation of damaged mitochondria by fusion and thus accelerates their deterioration.
Open questions
Even though mitochondria and their membrane poly peptide complexes have been studied intensely for more than than five decades, they remain a constant source of fascinating and unexpected new insights. Open questions abound, many of them of a fundamental nature and of direct relevance to homo wellness [61].
Apropos macromolecular structure and function, we do non however empathize the precise office of the highly conserved feature of ATP synthase dimers and dimer rows in the cristae and the interplay between the MICOS circuitous and the dimer rows in cristae germination. Are there other factors involved in determining crista size and shape?
Nosotros however do not know how complex I works, especially how electron transfer is coupled to proton translocation. What is the role of respiratory chain supercomplexes? Do they assist to prevent oxidative harm to mitochondria, and if so, how? And how does this touch on ageing and senescence?
We besides practice not know how the TIM and TOM protein translocases work, and what they wait like at high resolution. The aforementioned is true for the structure of the MICOS complex at the crista junctions. How does it ballast the cristae to the outer membrane, and how does information technology separate the cristae form the contiguous purlieus membrane? Similarly, the mechanisms of mitochondrial fission and fusion and the precise interest and coordination of the various protein complexes in this intricate procedure is a fascinating area of discovery.
The biogenesis and associates of large membrane protein complexes in mitochondria is largely unexplored. Where and exactly how do the respiratory chain complexes and the ATP synthase assemble? How is their assembly from mitochondrial and nuclear gene products coordinated? Does this involve feedback from the mitochondrion to the cytoplasm or the nucleus, and what is it?
And finally, how exactly are mitochondria implicated in ageing? Why do some cells and organisms live but for days, while others have lifespans of years or decades? Is this genetically programmed or simply a consequence of different levels of oxidative impairment? How is this damage prevented or controlled, and how does information technology affect the function of mitochondrial complexes? Is the breakdown of ATP synthase dimers also an effect of oxidative damage, and is it a cause of ageing?
It volition be challenging to detect answers to these questions because many of the protein complexes involved are thin, fragile and dynamic, and they do not lend themselves easily to well established methods, such as protein crystallography. Cryo-EM, which is currently undergoing rapid evolution in terms of high-resolution detail, will have a major impact just is limited to molecules above about 100 kDa [62]. Even better, more sensitive electron detectors than the ones that accept precipitated the recent resolution revolution, in combination with innovative image processing software, will yield more structures at higher resolution. However, small, rare and dynamic complexes will remain difficult to deal with. New labeling strategies in combination with other biophysical and genetic techniques are needed. Cloneable labels for electron microscopy, equivalent to green fluorescent protein in fluorescence microscopy, would be a dandy help; first steps in this management look promising [26]. In one case the structures and locations of the participating complexes take been determined, molecular dynamics simulations, which can analyze increasingly big systems, can help to empathise their molecular mechanisms. Without any doubt, mitochondria and their membrane protein complexes will remain an attractive research expanse in biology for many years to come.
References
-
Rizzuto R, De Stefani D, Raffaello A, Mammucari C. Mitochondria every bit sensors and regulators of calcium signalling. Nat Rev Mol Prison cell Biol. 2012;13(ix):566–78.
-
Pellegrino MW, Haynes CM. Mitophagy and the mitochondrial unfolded protein response in neurodegeneration and bacterial infection. BMC Biol. 2015;xiii:22.
-
Chandel NS. Mitochondria every bit signaling organelles. BMC Biol. 2014;12:34.
-
Bratic A, Larsson NG. The function of mitochondria in aging. J Clin Invest. 2013;123(iii):951–vii.
-
Gray MW, Lang BF, Cedergren R, Golding GB, Lemieux C, Sankoff D, et al. Genome structure and gene content in protist mitochondrial DNAs. Nucleic Acids Res. 1998;26(iv):865–78.
-
Amunts A, Brown A, Bai X, Llacer JL, Hussain T, Emsley P, et al. Construction of the yeast mitochondrial large ribosomal subunit. Scientific discipline. 2014;343(6178):1485–9.
-
Schmidt O, Pfanner N, Meisinger C. Mitochondrial poly peptide import: from proteomics to functional mechanisms. Nat Rev Mol Prison cell Biol. 2010;11(9):655–67.
-
Lackner LL. Shaping the dynamic mitochondrial network. BMC Biol. 2014;12:35.
-
Detmer SA, Chan DC. Functions and dysfunctions of mitochondrial dynamics. Nat Rev Mol Cell Biol. 2007;eight(11):870–9.
-
Hoppins South, Lackner Fifty, Nunnari J. The machines that split up and fuse mitochondria. Annu Rev Biochem. 2007;76:751–80.
-
Alexandre A, Reynafarje B, Lehninger AL. Stoichiometry of vectorial H+ movements coupled to electron transport and to ATP synthesis in mitochondria. Proc Natl Acad Sci U S A. 1978;75(eleven):5296–300.
-
Meeusen S, McCaffery JM, Nunnari J. Mitochondrial fusion intermediates revealed in vitro. Science. 2004;305:1747–52.
-
Gutman One thousand, Kotlyar AB, Borovok N, Nachliel E. Reaction of majority protons with a mitochondrial inner membrane grooming: fourth dimension-resolved measurements and their assay. Biochemistry. 1993;32(12):2942–vi.
-
Perkins GA, Frey TG. Recent structural insight into mitochondria gained by microscopy. Micron. 2000;31(one):97–111.
-
Liao M, Cao E, Julius D, Cheng Y. Construction of the TRPV1 ion aqueduct determined by electron cryo-microscopy. Nature. 2013;504(7478):107–12.
-
Kühlbrandt W. The resolution revolution. Science. 2014;343(6178):1443–four.
-
Davies KM, Strauss M, Daum B, Kief JH, Osiewacz HD, Rycovska A, et al. Macromolecular system of ATP synthase and complex I in whole mitochondria. Proc Natl Acad Sci U Due south A. 2011;108(34):14121–6.
-
India TA, Davey NE, Ulbrich P, Riches JD, de Marco A, Rumlova M, et al. Structure of the immature retroviral capsid at 8 A resolution by cryo-electron microscopy. Nature. 2012;487(7407):385–ix.
-
Bayrhuber M, Meins T, Habeck G, Becker S, Giller Thou, Villinger S, et al. Structure of the homo voltage-dependent anion aqueduct. Proc Natl Acad Sci U S A. 2008;105(forty):15370–5.
-
Llopis J, McCaffery JM, Miyawaki A, Farquhar MG, Tsien RY. Measurement of cytosolic, mitochondrial, and Golgi pH in single living cells with green fluorescent proteins. Proc Natl Acad Sci U S A. 1998;95(12):6803–viii.
-
Kukat C, Wurm CA, Spahr H, Falkenberg Thou, Larsson NG, Jakobs S. Super-resolution microscopy reveals that mammalian mitochondrial nucleoids take a uniform size and ofttimes incorporate a unmarried copy of mtDNA. Proc Natl Acad Sci U Due south A. 2011;108(33):13534–nine.
-
Kukat C, Davies K, Wurm CA, Spähr H, Bonekamp NA, Kühl I, et al. Cross-strand binding of TFAM to a single mtDNA molecule forms the mitochondrial nucleoid. Proc Natl Acad Sci U S A. 2015. In press.
-
Pfeffer Southward, Woellhaf MW, Herrmann JM, Forster F. Organization of the mitochondrial translation machinery studied in situ by cryoelectron tomography. Nat Commun. 2015;6:6019.
-
Schleyer M, Neupert W. Transport of proteins into mitochondria: translocational intermediates spanning contact sites between outer and inner membranes. Jail cell. 1985;43(ane):339–50.
-
Perkins Thou, Renken C, Martone ME, Young SJ, Ellisman One thousand, Frey T. Electron tomography of neuronal mitochondria: three-dimensional structure and organisation of cristae and membrane contacts. J Struct Biol. 1997;119(3):260–72.
-
Gold VA, Ieva R, Walter A, Pfanner North, van der Laan M, Kuhlbrandt W. Visualizing active membrane poly peptide complexes by electron cryotomography. Nat Commun. 2014;5:4129.
-
Pebay-Peyroula E, Dahout-Gonzalez C, Kahn R, Trezeguet V, Lauquin GJ, Brandolin Thou. Structure of mitochondrial ADP/ATP carrier in complex with carboxyatractyloside. Nature. 2003;426(6962):39–44.
-
Sjostrand FS. Electron microscopy of mitochondria and cytoplasmic double membranes. Nature. 1953;171(4340):30–2.
-
Palade GE. An electron microscope study of the mitochondrial structure. J Histochem Cytochem. 1953;1(four):188–211.
-
Davies KM, Anselmi C, Wittig I, Faraldo-Gomez JD, Kühlbrandt Due west. Structure of the yeast F1Fo-ATP synthase dimer and its role in shaping the mitochondrial cristae. Proc Natl Acad Sci U S A. 2012;109(34):13602–7.
-
Pfanner N, van der Laan Thou, Amati P, Capaldi RA, Caudy AA, Chacinska A, et al. Compatible nomenclature for the mitochondrial contact site and cristae organizing system. J Jail cell Biol. 2014;204(7):1083–vi.
-
von der Malsburg M, Muller JM, Bohnert M, Oeljeklaus South, Kwiatkowska P, Becker T, et al. Dual role of mitofilin in mitochondrial membrane organization and poly peptide biogenesis. Dev Cell. 2011;21(4):694–707.
-
Vogel F, Bornhovd C, Neupert W, Reichert Every bit. Dynamic subcompartmentalization of the mitochondrial inner membrane. J Cell Biol. 2006;175(2):237–47.
-
von Ballmoos C, Wiedenmann A, Dimroth P. Essentials for ATP synthesis by F1F0 ATP synthases. Annu Rev Biochem. 2009;78:649–72.
-
Watt IN, Montgomery MG, Runswick MJ, Leslie AG, Walker JE. Bioenergetic toll of making an adenosine triphosphate molecule in animal mitochondria. Proc Natl Acad Sci U S A. 2010;107(39):16823–7.
-
Stock D, Leslie AGW, Walker JE. Molecular architecture of the rotary motor in ATP synthase. Scientific discipline. 1999;286:17701705.
-
Allen RD, Schroeder CC, Fok AK. An investigation of mitochondrial inner membranes by rapid-freeze deep-etch techniques. J Jail cell Biol. 1989;108(half-dozen):2233–twoscore.
-
Strauss M, Hofhaus G, Schroder RR, Kühlbrandt W. Dimer ribbons of ATP synthase shape the inner mitochondrial membrane. EMBO J. 2008;27(seven):1154–lx.
-
Allegretti One thousand, Klusch N, Mills DJ, Vonck J, Kühlbrandt W, Davies KM. Horizontal membrane-intrinsic alpha-helices in the stator a-subunit of an F-blazon ATP synthase. Nature. 2015;521(7551):237–forty.
-
Kühlbrandt Westward, Karen D. ATP synthases: A new twist for an aboriginal motorcar. Trends Biochem Sci. 2015. In press.
-
Paumard P, Vaillier J, Coulary B, Schaeffer J, Soubannier V, Mueller DM, et al. The ATP synthase is involved in generating mitochondrial cristae morphology. EMBO J. 2002;21(iii):221–30.
-
Bornhövd C, Vogel F, Neupert West, Reichert Equally. Mitochondrial membrane potential is dependent on the oligomeric state of F1F0-ATP synthase supracomplexes. J Biol Chem. 2006;281(20):13990–viii.
-
Giorgio Five, von Stockum S, Antoniel M, Fabbro A, Fogolari F, Forte G, et al. Dimers of mitochondrial ATP synthase form the permeability transition pore. Proc Natl Acad Sci U S A. 2013;110(fifteen):5887–92.
-
Hunte C, Koepke J, Lange C, Roßmanith T, Michel H. Construction at 2.3 Å resolution of the cytochrome bc1 complex from the yeast Saccharomyces cerevisiae co-crystallized with an antibody Fv fragment. Structure. 2000;eight:669–84.
-
Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, et al. The whole structure of the xiii-subunit oxidized cytochrome c oxidase at 2.8 A. Scientific discipline. 1996;272(5265):1136–44.
-
Zickermann Five, Wirth C, Nasiri H, Siegmund Yard, Schwalbe H, Hunte C, et al. Mechanistic insight from the crystal structure of mitochondrial complex I. Scientific discipline. 2015;347(6217):44–9.
-
Efremov RG, Baradaran R, Sazanov LA. The compages of respiratory complex I. Nature. 2010;465(7297):441–5.
-
Vinothkumar KR, Zhu J, Hirst J. Architecture of mammalian respiratory complex I. Nature. 2014;515(7525):80–4.
-
Schägger H, Pfeiffer Yard. Supercomplexes in the respiratory bondage of yeast and mammalian mitochondria. EMBO J. 2000;19(8):1777–83.
-
Schäfer East, Seelert H, Reifschneider NH, Krause F, Dencher NA, Vonck J. Compages of active mammalian respiratory concatenation supercomplexes. J Biol Chem. 2006;281(22):15370–five.
-
Althoff T, Mills DJ, Popot JL, Kühlbrandt Due west. Arrangement of electron send chain components in bovine mitochondrial supercomplex I1III2IV1. EMBO J. 2011;30(22):4652–64.
-
Lapuente-Brun Due east, Moreno-Loshuertos R, Acin-Perez R, Latorre-Pellicer A, Colas C, Balsa E, et al. Supercomplex assembly determines electron flux in the mitochondrial electron ship chain. Science. 2013;340(6140):1567–seventy.
-
Mileykovskaya E, Penczek PA, Fang J, Mallampalli VK, Sparagna GC, Dowhan W. Arrangement of the respiratory chain complexes in Saccharomyces cerevisiae supercomplex III2IV2 revealed by unmarried particle cryo-electron microscopy. J Biol Chem. 2012;287(27):23095–103.
-
Blaza JN, Serreli R, Jones AJ, Mohammed Thousand, Hirst J. Kinetic evidence confronting partition of the ubiquinone pool and the catalytic relevance of respiratory-chain supercomplexes. Proc Natl Acad Sci U Due south A. 2014;111(44):15735–40.
-
Li P, Nijhawan D, Budihardjo I, Srinivasula SM, Ahmad M, Alnemri ES, et al. Cytochrome c and dATP-dependent formation of Apaf-one/caspase-9 complex initiates an apoptotic protease cascade. Cell. 1997;91(4):479–89.
-
Daum B, Walter A, Horst A, Osiewacz HD, Kühlbrandt W. Age-dependent dissociation of ATP synthase dimers and loss of inner-membrane cristae in mitochondria. Proc Natl Acad Sci U South A. 2013;110(38):15301–six.
-
Navarro A, Boveris A. The mitochondrial energy transduction organization and the aging process. Am J Physiol Cell Physiol. 2007;292(2):C670–86.
-
Dröse S, Brandt U. Molecular mechanisms of superoxide production by the mitochondrial respiratory concatenation. Adv Exp Med Biol. 2012;748:145–69.
-
Twig Chiliad, Shirihai Os. The interplay betwixt mitochondrial dynamics and mitophagy. Antioxid Redox Bespeak. 2011;14(10):1939–51.
-
Scheckhuber CQ, Erjavec N, Tinazli A, Hamann A, Nyström T, Osiewacz Hard disk drive. Reducing mitochondrial fission results in increased life span and fitness of two fungal ageing models. Nat Cell Biol. 2007;9(1):99–105.
-
Calvo SE, Mootha VK. The mitochondrial proteome and human illness. Annu Rev Genomics Hum Genet. 2010;11:25–44.
-
Henderson R. The potential and limitations of neutrons, electrons and X-rays for diminutive resolution microscopy of unstained biological molecules. Q Rev Biophys. 1995;28:171–93.
-
Rees DM, Leslie AG, Walker JE. The construction of the membrane extrinsic region of bovine ATP synthase. Proc Natl Acad Sci U S A. 2009;106(51):21597–601.
-
Symersky J, Pagadala 5, Osowski D, Krah A, Meier T, Faraldo-Gomez JD, et al. Construction of the c(10) ring of the yeast mitochondrial ATP synthase in the open conformation. Nat Struct Mol Biol. 2012;19(5):485–91. S481.
Acknowledgements
I thank Bertram Daum, Karen Davies, Vicki Aureate, Klaus Pfanner and Thomas Langer for helpful comments on the manuscript.
Author information
Affiliations
Respective writer
Additional files
Boosted file i:
Movie S1. Mitochondria in a human endothelial cell. Fourth dimension-lapse motion picture of the dynamic mitochondrial network stained with a fluorescent dye. Long filamentous mitochondria occasionally undergo fission, while smaller parts of the network fuse into longer tubes. (Courtesy of Jürgen Bereiter-Hahn, Goethe University, Frankfurt, Germany) (AVI 137694 kb)
Additional file 2:
Pic S2. Dimer rows of mitochondrial ATP synthase in cristae membranes. The iii-dimensional volume of a pocket-size P. anserina mitochondrion obtained by cryo-ET shows that ATP synthase dimers grade long rows forth cristae ridges. The outer membrane is grey, the inner membrane and cristae membranes are calorie-free blueish. The Fi heads of the ATP synthase are indicated in yellow. (Adapted from [17]) (MP4 7740 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give advisable credit to the original author(s) and the source, provide a link to the Artistic Commons license, and indicate if changes were made. The Artistic Eatables Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/i.0/) applies to the data made available in this article, unless otherwise stated.
Reprints and Permissions
Virtually this article
Cite this commodity
Kühlbrandt, Westward. Structure and function of mitochondrial membrane protein complexes. BMC Biol thirteen, 89 (2015). https://doi.org/10.1186/s12915-015-0201-10
-
Published:
-
DOI : https://doi.org/ten.1186/s12915-015-0201-ten
Keywords
- Respiratory Concatenation Complex
- Mitochondrial Fission
- Membrane Protein Complex
- Boundary Membrane
- Bovine Heart Mitochondrion
Source: https://bmcbiol.biomedcentral.com/articles/10.1186/s12915-015-0201-x
0 Response to "what must be true about the inner mitochondrial membrane for the gradient to work"
Post a Comment